How do small sugars bind to each other?
Broadband rotational spectroscopy reveals the precise structure of the glycolaldehyde dimer
Sugars are important building blocks in nature. They can occur on the cell surface, where they interact with other cells in a very characteristic way, as for example found in the synthesis of the blood group antigens A and B. Both antigens are built up by a long chain of sugar molecules and only differ by a selectively bound terminal sugar. On the molecular level, the way how molecules selectively bind to each other is controlled by the interplay of different intra- and intermolecular interactions.
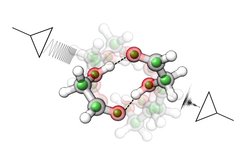
Depending on their structure, some parts of the molecules might attract and some might repel each other. As a result, the formation of molecular complexes can be highly selective and is also described as molecular recognition. To understand such molecular recognition processes on the molecular level, the researchers investigated the smallest sugar, glycolaldehyde. It is also the first and so far only sugar detected in space, and an important player in the discussion on the origin of life in space. The technique the researchers employed is broadband rotational spectroscopy in the gas phase. It is an ideal tool to precisely determine the structure of molecules and molecular complexes.
“The aim of this study was to determine in detail how two sugar molecules bind to each other,” says PhD student Sabrina Zinn, first author of this work. “This is not intuitive because sugars offer many different binding sites so that a variety of outcomes can be envisioned.” Indeed, high-resolution broadband rotational spectroscopy reveals two different ways how the glycolaldehyde molecules bind to each other. “The precise structures that we obtained allow us to identify the dominant interaction mechanisms between the sugar molecules. It will now be interesting to see how these findings can be extended to larger sugars,” concludes group leader Melanie Schnell.






